Advances in dairy Research
Open Access
ISSN: 2329-888X
ISSN: 2329-888X
Research Article - (2019)Volume 7, Issue 3
Background: Fiore Sardo PDO is one of the oldest Mediterranean hard cheeses, exclusively produced in Sardinia (Italy) from raw whole ewe’s milk. Some manufacturers, improperly, submit raw milk to heat-treatment. Aim of this study was to evaluate the proteomic profile of Fiore Sardo PDO and to investigate possible differences between cheese made from raw milk and from heat-treated milk.
Materials and methods: Starting from the same ovine bulk milk, eight cheese wheels of Fiore Sardo PDO were produced, four from raw milk and four from heat-treated milk. The subsequent production steps were the same for both types of cheese as was sampling at different ripening times. Samples were analyzed by Urea-PAGE electrophoresis. Afterwards, blind analysis of 32 cheese samples, produced by 17 different cheesemakers was performed to evaluate the method’s analytical performance.
Results: Urea-PAGE analysis showed the presence of a protein band only in cheese made from raw milk, regardless of cheese ripening time. Protein identification analysis by liquid chromatography mass spectrometry identified two different proteins in the band, alpha-S1 and alpha-S2 casein. Blind analyses conducted to verify the analytical performance of the method showed that it could be a useful tool for the protection of this typical agri-food product.
Conclusion: We developed a simple, robust, and economic method for discriminating between Fiore Sardo PDO cheese made from raw milk with up to 24 months of maturation and Sardinian sheep milk cheese made from heattreated milk.
Urea-PAGE; Fiore Sardo PDO; Heat-treated milk; Food fraud; Food authenticity
Fiore Sardo PDO is one of the oldest known Mediterranean hard cheeses, with origins dating back to the Bronze Age. It is produced in Sardinia (Italy) primarily by artisanal farms from raw whole ewe’s milk curdled using lamb rennet. The rind varies in color from deep yellow to dark brown and encases a paste that varies from white to straw-yellow. The sharpness of the flavor depends on the length of maturation. Young Fiore Sardo is about 4 months old, while the ripened type is matured for more than 6 months up to over 2 years.
It was awarded Protected Designation of Origin (PDO) status by the European Commission [1].
The Fiore Sardo PDO cheesemaking protocol entails milk curdling with lamb rennet paste at 34°C to 36°C. The curd is cut and left to drain. As soon as the rinds are formed, they are superficially treated with warm water or scotta (a by-product of ricotta cheese production) to develop a thick outer rind and are then salted. Cheese maturation includes smoking treatment for two weeks at 18°C to 20°C, ripening in a cellar at 10°C to 15°C and final treatment of the rinds with an emulsion of olive oil, salt, and wine vinegar.
In the last twenty years, many studies have been done to characterize both sheep milk and Fiore Sardo PDO. Regarding milk, no immediate parallels can be drawn between sheep and bovine milk, due to differences in lipid and protein concentrations, therefore the alkaline phosphatase used for bovine milk is not an efficient marker in ovine milk but the enzymatic activities of α-L-fucosidase and γ- glutamyltransferase have been studied and can be a reliable markers of thermization and pasteurization of sheep milk [2]. Many aspects of the Fiore Sardo PDO have been investigated such as microbiological, chemical, sensorial characteristics [3-5], evaluation of biogenic amines [6] and studies on the use of lamb rennet and the relationship between its enzymatic composition and the proteolytic and lipid pattern of the Fiore Sardo PDO [7,8].
The use of raw milk is one of the most distinctive characteristics of Fiore Sardo PDO, while other Sardinian sheep milk cheeses, Pecorino Sardo PDO and Pecorino Romano PDO, can be produced from heat-treated milk. Some manufacturers improperly heattreat all the raw milk (68°C for 15 seconds) or pasteurize it (72°C for 15 seconds) before starting cheese productions, also for the manufacture of Fiore Sardo PDO. Indeed, such treatment ensures hygiene of the industrial production lines, increases product safety and yield of the product, in addition to prolonging average shelf life and preventing the appearance of possible off-flavors [9].
Discriminating between dairy products manufactured from raw or heat-treated milk is an analytical challenge. Previous studies on different cheeses have evaluated changes in enzymatic activity (lactosylation) of caseins [10,11]. A study using Magnetic Resonance Imaging (MRI) successfully discriminated between Fiore Sardo PDO produced according to specifications and non-genuine cheese [12] and, more recently, promising results were obtained with gas chromatography-mass spectrometry approach [13].
Chromatographic and electrophoretic methods are useful tools to characterize cheeses also in relation to ripening time [14]. Furthermore, both processing and heat treatment influence the structure and composition of milk proteins and the characteristics of cheeses [15-17]. For this study we have focused on proteins to evaluate the proteomic profile of Fiore Sardo PDO and to determine possible differences between cheese made from raw milk and from heat-treated milk.
Samples and experimental design
Our study was conducted in two steps: first to develop and optimize the method, and second to evaluate its analytical performance. The analyses were started on eight cheese wheels of Fiore Sardo PDO, prepared and certified by an artisanal manufacturer following the technical specifications for the production using 210 liters of ovine bulk milk, divided into two equal parts. Four samples were made from Raw milk (R) (as required by the original recipe) and four samples from the same milk after Heat-Treatment (H-T) (68°C for 15 seconds). All subsequent production steps were the same for both types (R/H-T) of cheese, as briefly described below. Milk in each vat (105 liters) was heated up to 35°C and autochthonous adjunct cultures were inoculated. After 10 min, the milk is coagulated using lamb rennet paste in 20-30 min at 33-35°C, then the curd is cut and crumbled manually and kept in the vat for a few minutes. The curd is cut by hand into small pieces, the size of a rice grain, and whey is removed by hand pressing before moulding. The curd is then transferred to moulds that give the cheese its typical shape and put in warm room up to reach optimal pH. The cheeses are brine salted for 36 h; afterwards, they are placed on a bed of straw and slightly smoked for 12 days using wood of Mediterranean plants (Arbutus unedo L., Pistacia Ientiscus L., Mirtus communis L., Phillyrea latifolia L., Quercus var. pubescens Willd., Quercus IIex L, Quercus suber L., Olea europea var. sylvestris Brot.). The temperature and relative humidity of maturation can vary between 15°C and 18°C and 80% and 85%, respectively. During maturation, the cheese is turned and rubbed with olive oil to avoid dehydration of the rind.
In order to preserve all the characteristics of Fiore Sardo, cheese wheels were kept in the cellars and periodically sent to the laboratory for sampling and analysis. Sampling was performed at different ripening times (141, 183, 197, 294, 394, and 519 days) for both types (R/H-T) of cheese. All analytical samples, cut from the inner or intermediate part of the cheese wheel, avoiding the rind, were stored at 4°C until analysis.
Blind analysis of 32 cheese samples, produced by 17 different cheesemakers was performed to evaluate the method’s analytical performance. Samples were taken at different ripening stages, between 7 and 25 months (Table 1). The analyses were repeated by two technicians to ensure repeatability of the results.
| Sample | Manufacturer | Manufacturer certification | Production date | Ripening stage (month) | Presence of the protein band | Urea-PAGE Results |
|---|---|---|---|---|---|---|
| ID | ||||||
| 1 | A | R | 1/2/2018 | 11 | YES | R |
| 2 | B | R | 1/1/2018 | 12 | YES | R |
| 3 | B | R | 1/6/2018 | 7 | YES | R |
| 4 | B | R | 1/3/2017 | 22 | n.e.* | n.e.* |
| 5 | C | R | 1/2/2018 | 11 | YES | R |
| 6 | C | R | 1/5/2018 | 8 | YES | R |
| 7 | D | H-T | 1/1/2016 | 24 | NO | H-T |
| 8 | D | H-T | 1/1/2016 | 24 | NO | H-T |
| 9 | D | R | 1/3/2018 | 10 | YES | R |
| 10 | E | R | 1/3/2018 | 10 | YES | R |
| 11 | F | R | 11/12/2016 | 14 | YES | R |
| 12 | F | R | 9/2/2017 | 12 | YES | R |
| 13 | G | R | 21/12/2016 | 13 | YES | R |
| 14 | H | R | 31/01/2016 | 24 | YES | R |
| 15 | B | R | 31/12/2015 | 25 | n.e.* | n.e.* |
| 16 | I | R | 12/4/2017 | 10 | YES | R |
| 17 | L | R | 1/1/2017 | 13 | YES | R |
| 18 | M | R | 15/05/2017 | 9 | YES | R |
| 19 | N | R | 31/01/2017 | 12 | YES | R |
| 20 | D | R | 30/04/2017 | 9 | YES | R |
| 21 | M | R | 12/12/2016 | 14 | YES | R |
| 22 | O | R | 28/02/2017 | 11 | YES | R |
| 23 | I | R | 22/12/2016 | 13 | YES | R |
| 24 | N | R | 31/03/2017 | 10 | YES | R |
| 25 | H | R | 31/03/2017 | 10 | YES | R |
| 26 | P | R | 31/01/2017 | 12 | YES | R |
| 27 | L | R | 31/05/2017 | 8 | YES | R |
| 28 | Q | R | 30/06/2017 | 7 | YES | R |
| 29 | R | R | 10/12/2016 | 14 | YES | R |
| 30 | S | R | 11/12/2016 | 14 | YES | R |
| 31 | B | R | 30/04/2016 | 21 | n.e.* | n.e.* |
| 32 | B | R | 30/04/2017 | 9 | YES | R |
n.e.*: Not
Available
Table 1: List of the 32 cheese samples used for the blind analysis, ID for the 17 different manufacturers, Manufacturer certification, production date, ripening stage at analysis (expressed in months), presence of the protein band at Urea-PAGE analysis, and sample type (raw [R] or heat-treated [H-T] milk).
Protein extraction
Finely grated cheese (2 g) and 0.5 ml β-mercaptoethanol were dispersed in 10 ml of a buffer solution (0.06 M Tris, 8 M urea, pH 6.8). The mixture was held at 40°C for 45 min with periodic vigorous stirring and then filtered through glass wool. Prior to loading them onto gels, the filtrates were mixed in a 1:1 ratio with a marker dye solution consisting of 0.06 M Tris at pH 6.8, 8 M urea, 0.02% bromophenol blue, and 20% glycerin. For all samples, a volume of 10 μL, corresponding to ̴ 0.4 mg of total protein content, as quantified by Qubit Protein Assay (Thermo Fisher Scientific, Waltham, MA, USA), was applied to each well in the gel.
If not analyzed immediately, the protein extracts were stored at -20°C. Before the analysis, the extracts were thawed for at least 1 hour at Room Temperature (RT), incubated for 20 min at 45°C, and then mixed with the marker dye solution.
For each extraction session during the second step of analyses, a sample made from raw milk and one made from heat-treated milk from those prepared and certified for the first step were re-extracted and loaded onto gels as controls.
Urea-PAGE analysis
Urea-PAGE analysis was carried out on running gel (20 × 20 cm, 1.5 mm thick) consisting of T=12%, C=2.7%, pH 8.8 (buffered by 1.875 M Tris) and 3.2 M urea. The stacking gel solution consisted of T=8.5%, C=2.7%, pH 6.8 (buffered by 0.5 M Tris) and 2.24 M urea. The buffer for Urea-PAGE consisted of 12 g Tris and 57.4 g glycine in 4 L solution and the electrophoresis was carried out at 200 V for 40 min followed by 240 min at 280 V.
Staining
After electrophoresis, the gels were fixed in a solution containing 40% methanol and 10% acetic acid for 1 hour at RT, washed 4 times for 15 min each in ultrapure water, then stained with Blue Silver [18]. The gels were scanned with a GS-900™ Calibrated Densitometer and Image Lab software (Bio-Rad, Hercules, CA, USA) and analyzed to detect differences between cheese made from raw milk and from heat-treated milk. The bands of interest were excised and analyzed by mass spectrometry as described below.
Mass spectrometry
Protein bands from Urea-PAGE were cut out and the proteins were in-gel digested with trypsin (Sigma-Aldrich, Saint Louis, MO, USA). The peptide digests were desalted by a Discovery® DSC-18 Solid Phase Extraction (SPE) 96-well plates (25 mg/well) (Sigma-Aldrich.) and then vacuum evaporated prior to Liquid Chromatography Mass Spectrometry (LC-MS) analysis.
The digested samples were analyzed on a micro-LC (Eksigent Technologies, Dublin, CA, USA) interfaced to a 5600+ TripleTOF (TOF= time of flight) mass spectrometer system (AB SCIEX, Concord, Canada) equipped with a DuoSpray ion source and a CDS (Calibrant Delivery System). The LC column was a Halo Fused C18. The mobile phase was a mixture of 0.1% (v/v) formic acid in water (A) and 0.1% (v/v) formic acid in acetonitrile (B), eluting at a flow rate of 15.0 μL min−1 at an increasing concentration of solvent B from 2 to 40% in 30 min. The injection volume was 4.0 μL, and the oven temperature was set at 40°C. For identification purposes, MS analysis was performed at a mass range of 100–1500 Da (Time of Flight [TOF] scan with an accumulation time 0.25 s), followed by a MS/MS product ion scan from 200 to 1250 Da (accumulation time 5.0 ms) with the abundance threshold set at 30 cps (35 candidate ions can be monitored during a cycle). The ion source parameters in electrospray positive mode were set as follows: curtain gas (N2) at 2.736939 Bar, nebulizer gas GAS1 (N2) at 2.736939 Bar and GAS2 (N2) at 2.392201 Bar, ion spray floating voltage (ISFV) at 5000 V, source temperature at 450°C, and declustering potential at 25 V. The MS data were acquired with Analyst TF 1.7 (AB SCIEX) and the files were searched using Mascot v. 2.4 (Matrix Science Inc., Boston, MA, USA). Trypsin as digestion enzyme and one missed cleavage were specified. The instrument was set to ESI-QUAD-TOF (ESI=Electro Spray Ionization), and the following modifications were specified for the search: carbamidomethyl cysteins as fixed modification and oxidized methionine phosphorylated serine/threonine and phosphorylated tyrosine as variable modification. A search tolerance of 0.08 Da was specified for the peptide mass tolerance and 10 ppm for the MS/MS tolerance. The peptide charges to search for were set to 2+, 3+, and 4+, and the search was set on monoisotopic mass. The UniProt Swiss-Prot Ovis aries reviewed database (vers. 28/03/2107, containing 693 sequence entries) was used.
Urea-PAGE analysis highlighted the presence of a protein band only in the cheese wheels made from raw milk, regardless of cheese ripening time. Since Fiore Sardo PDO can be sold starting from 105 days of maturation, we tested the samples especially produced for this study from 141 to 519 days and over. The difference in protein band profile was found to remain stable over time (Figure 1).
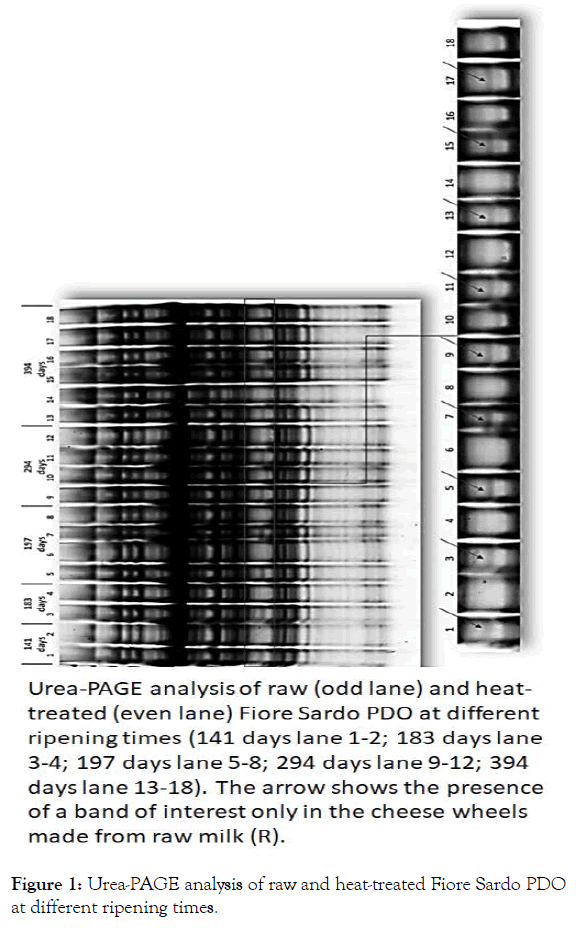
Figure 1: Urea-PAGE analysis of raw and heat-treated Fiore Sardo PDO at different ripening times.
At the beginning of our study, starting from a well-described protocol [14], different parameters were reviewed to improve the method’s analytical performance. Compared to the initial protocol, which involved the use of 0.2 g of grated cheese, we increased the quantity up to 2 g to enhance visibility of the band of interest. The total protein quantifications obtained by the Qubit method and the loading of serial dilutions, conducted on several samples from both groups, and enabled us to determine that the band of interest, if present, remained visible by loading a few micrograms of total proteins (data not shown). The detection limit is at 0.15 mg (total protein content), which is decidedly lower than the concentration normally loaded in each well (0.4 mg) during the later analyses. The use of Blue Silver [18], a more sensitive staining than Coomassie R-250, allowed us to obtain protein bands with more intense signals. Different sampling sites, starting from the center of the cheese wheel, were initially used (inner, intermediate and rind area), but the rind area should be avoided since it is richer in salts, which can interfere with the electrophoretic run, and it is more exposed to the external environment.
The band of interest was excised from the gel and protein identification analysis was performed by liquid chromatography mass spectrometry (LC-MS/MS). LC-MS/MS analysis identified two proteins in the band, alpha-S1 and alpha-S2 casein. Table 2 presents the two proteins with a Mascot score of 1632 and 2489, respectively, for alpha-S1 and alpha-S2 casein.
| Protein (Mascot Score) | Sequence | Nominal Mass (Da)/Calculated pI | Sequence Coverage | Uniprot Accession Number |
|---|---|---|---|---|
| Alpha-S1-casein (1632) | MKLLILTCLVAVALARPKHPIKHQGLSSEVLNENLLRFVVAPFPEVFRKENINELSKDIGSESIEDQAMEDAKQMKAGSSSSSEEIVPNSAEQKYIQKEDVPSERYLGYLEQLLRLKKYNVPQLEIVPKSAEEQLHSMKEGNPAHQKQPMIAVNQELAYFYPQLFRQFYQLDAYPSGAWYYLPLGTQYTDAPSFSDIPNPIGSENSGKITMPLW | 24374/5.32 | 39% | P04653 (CASA1_SHEEP) |
| Alpha-S2-casein (2489) | MKFFIFTCLLAVALAKHKMEHVSSSEEPINISQEIYKQEKNMAIHPRKEKLCTTSCEEVVRNADEEEYSIRSSSEESAEVAPEEVKITVDDKHYQKALNEINQFYQKFPQYLQYLYQGPIVLNPWDQVKRNAGPFTPTVNREQLSTSEENSKKTIDMESTEVFTKKTKLTEEEKNRLNFLKKISQYYQKFAWPQYLKTVDQHQKAMKPWTQPKTNAIPYVRYL | 26486/7.66 | 52% | P04654 (CASA2_SHEEP) |
Table 2: Proteins identified with Mascot score (first column), peptide sequences (in bold) obtained by μLC-MS (second column), calculated molecular mass and isoelectric point (third column), sequence coverage (fourth column), and accession number (fifth column). LC-MS/MS results for protein identification show the presence of both Alpha-S1 and Alpha-S2 casein.
Blind analyses were carried out for confirming the results and evaluating the method’s analytical performance. They gave consistent results when were repeated by two different technicians. The two samples produced with heat-treated milk were correctly identified (Table 1 and Figure 2). None of the samples made from raw milk was erroneously identified as made from heat-treated milk. Only 3 out of 30 samples (nos. 4-15-31) were unverifiable because they showed a protein profile slightly different from the others. An interesting result emerged at the end of the analyses, when we asked for information about the type of milk (R or H-T) used and about the different manufacturers. The 32 samples came from 17 different producers, but the three unverifiable samples (nos. 4-15-31) came from the same cheese maker and were ripened for 22, 25, and 21 months, respectively. Three other samples (nos. 2-3-32) from the same manufacturer, ripened for 12, 7, and 9 months respectively, were correctly identified.

Figure 2: Urea-PAGE blinded analysis.
We assume that there was a factor that influenced the protein profile during long-term ripening (>21 months) for this specific producer, linked to the conditions (temperature/humidity) of maturation in its cellar. Fiore Sardo PDO made by other producers with up to 24 months of maturation have been correctly recognized, and investigating the conditions of the cellars we knew that, in those correctly identified cheeses, there is an aeration system that allows maintaining more controlled conditions. The fact that, in some particular samples, the profile showed differences in its ensemble should not be considered a limiting factor, because all cheese samples made from raw milk and all samples made from heattreated milk were correctly identified. Certainly, other factors are important and must be considered such as seasonality, feeding and use of lamb rennet paste. Cheeses were produced at different times through the year, thus in different moments of ewe’s lactation cycle, and come from farms located in different municipalities of Sardinia. All the cheese-makers used lamb rennet paste, and this is crucial because it contain lipolytic enzymes that hydrolyze milk fat triglycerides yelding free fatty acids, which impart the characteristic piquant taste to cheese [7].
Food fraud and authenticity are a growing problem with a global impact on all steps in the food chain. Since consumers demand safe and certified foods that meet the characteristics claimed on the label, there is an increasing need to implement measures to ensure it. Unfortunately, because of the complexity involved, instances of fraud within the food supply chain go undetected [19,20]. In the case of Fiore Sardo PDO, in addition to the consumer, the artisanal producers who honestly produce cheese according to the original recipe and technical specifications must be protected. The use of heat-treated milk to produce non-genuine Fiore Sardo and the sale of cheese that does not comply with the PDO regulations constitute deception of the consumer and major economic damage for small farmers and the local economy.
The need to distinguish unequivocally and objectively between cheeses made from raw milk and those made from heat-treated milk is shared by many PDO cheeses: Parmigiano Reggiano, Grana Padano, Castelmagno, and Fontina are just a few examples of Italian PDO cheeses manufactured starting from raw milk. Although the influence of heat treatment on the composition and microbiological quality of raw milk has been investigated [21,22] few methods are available that can distinguish between cheeses produced from raw or heat-treated milk. Finding a rapid and reliable analytical method would therefore be valuable for both scientific and technological reasons [23].
Developing and optimizing analytical methods based on various different approaches can improve the chance of detecting anomalies and frauds. In this effort, proteomic approaches are useful for food product characterization and authentication and thus provide a reliable tool to uncover fraudulent practices [24–28] and, concerning Fiore Sardo PDO, a multisciplinary approach could be planned analyzing samples both with UREA-PAGE and other effective methods such as NMR and GC-MS.
The use of raw milk is one of the most distinctive characteristics of Fiore Sardo PDO while other Sardinian sheep milk cheeses, Pecorino Sardo PDO and Pecorino Romano PDO, can be made from heat-treated milk. There is a need for a method that can discriminate between genuine Fiore Sardo PDO, produced according to PDO specifications, and non-genuine cheese. With this study we developed a simple, robust, and economic method for discriminating between Fiore Sardo PDO cheese made from raw milk with up to 24 months of maturation and Sardinian sheep milk cheese made from heat-treated milk.
The authors are grateful to the artisanal cheese manufacturers for their collaboration and for providing the cheese samples.
This research was funded by the Italian Ministry of Health [grant number IZS PLV 10/13RC].
None to report.
Citation: Mazza M, Guglielmetti C, Brusadore S, Sciuto S, Esposito G, Caramelli M, et al. (2019) A Proteomic Approach to the Safeguard of a Typical Agri-Food Product: Fiore Sardo PDO. Adv Dairy Res 7:228. doi: 10.35248/2329-888X.19.7.228
Received: 27-Sep-2019 Accepted: 12-Oct-2019 Published: 18-Oct-2019 , DOI: 10.35248/2329-888X.19.7.228
Copyright: © 2019 Mazza M, et al. This is an open access article distributed under the term of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.