
Journal of Clinical and Cellular Immunology
Open Access
ISSN: 2155-9899

ISSN: 2155-9899
Short Communication - (2022)Volume 13, Issue 5
Aplastic Anemia (AA) is a hematopoietic failure symptom of the bone marrow that can be caused by a variety of factors such as T-cell mediated immune attack, genetic-related somatic mutations, cytogenetic abnormalities, and defective telomerase functions. Recently, Single-cell RNA-sequencing (scRNA-seq) analysis of bone marrow-derived Hematopoietic Stem and Progenitor Cells (HSPCs) from AA patients also revealed the involvement of selective lineage disruption, altered alternative splicing, and polyadenylation in AA pathogenesis . However, these new mechanisms did not provide any new therapeutic approaches for AA, as novel therapies are urgently needed for patients who received no durable benefit from immunosuppressive therapy combined with the Thrombopoietin Receptor (TpoR) activator and patients who underwent allogeneic hematopoietic stem cell transplantation. Therefore, further research is warranted for a deeper understanding of AA and better treatment of this disease.
Anemia; Immunosuppressive; T-cell; Immune attack
Autophagy plays an important role in removing activated mitochondria and controlling oxidative metabolism, thereby maintaining HSPCs stemness and regenerative potential [1-7]. In addition, autophagy also affects T cells in many aspects [8]. For example, autophagy in T cells can directly promote T cell expansion, survival, and differentiation, or indirectly enhance T cell antigen recognition via modulating antigen-presenting cells [9,10]. Previous studies have shown that spermidine, a natural polyamine compound, participates in many physiological processes that regulate autophagy, including the inhibition of several acetyltransferases, such as EP300 [11,12], activating the AKT/AMPK signal pathway, and inhibiting the mTOR pathway[13]. Spermidine can trigger FoxO transcription through AKT dephosphorylation, AMPK phosphorylation, and mTOR inactivation, which in turn increases ATGS and inhibits acetyltransferase EP300 expression. Consequently, the binding of ATGS and EP300 to P62 and LC3, two crucial autophagy proteins, enhances autophagy flux. Given that AA is an autoreactive T cell-mediated bone marrow HSPC disorder, it is speculative whether autophagy and spermidine affect the occurrence of AA.
To test the above hypothesis, we collected samples from 38 patients with idiopathic aplastic anemia and 5 healthy donors (Supplementary Table S1), including bone marrow, bone marrow supernatant, peripheral blood, and peripheral plasma. Bone marrow supernatant samples were subjected to Ultra-High Performance Liquid Chromatography−Tandem Mass Spectrometry (UHPLC-MS /MS) analysis.
The raw sequence data reported in this paper have been deposited in the Genome Sequence Archive (Genomics, Proteomics & Bioinformatics 2021) in National Genomics Data Center (Nucleic Acids Res 2021), China National Center for Bioinformation/Beijing Institute of Genomics, Chinese Academy of Sciences (GSA: HRA003557) that are publicly accessible at https://ngdc.cncb.ac.cn/gsa.
We detected a significant decrease of spermidine and its related metabolites in the metabolomics data sets, including spermine and glutathione, in AA samples (Figure 2). The Area Under the ROC Curves (AUC) of the spermine, spermidine, and glutathione were 0.975, 0.950, and 0.988, respectively. These results suggested that spermidine and its related metabolites may have a diagnostic potential for AA (Figure 3). To further investigate whether spermidine regulates autophagy levels in AA, we performed transmission electron microscopy and observed that the number of autophagic vacuoles decreased in AA samples compared to that in healthy donors (Figure 4). The decrease of autophagic vacuoles, together with the decrease of LC3B and increase of p62, suggested that the autophagic degradation process, also known as autophagy flux, was blocked in AA patients (Figures 3 and 5). In accordance, we found that exogenously added spermidine rescued autophagic flux in cultured bone marrow cells in vitro (Figure 1).
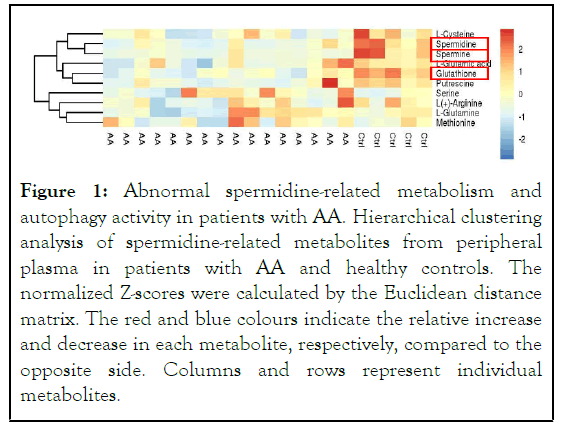
Figure 1: Abnormal spermidine-related metabolism and autophagy activity in patients with AA. Hierarchical clustering analysis of spermidine-related metabolites from peripheral plasma in patients with AA and healthy controls. The normalized Z-scores were calculated by the Euclidean distance matrix. The red and blue colours indicate the relative increase and decrease in each metabolite, respectively, compared to the opposite side. Columns and rows represent individual metabolites.
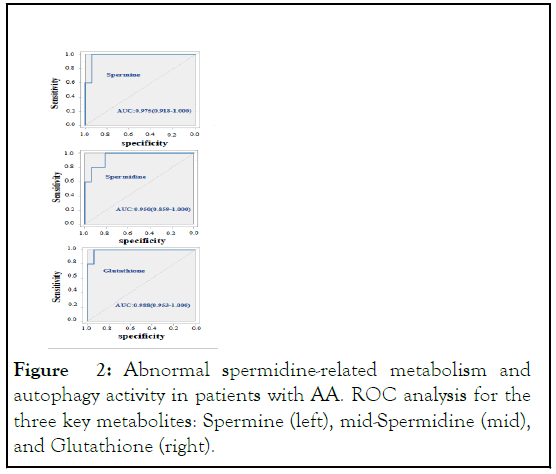

Figure 2: Abnormal spermidine-related metabolism and autophagy activity in patients with AA. ROC analysis for the three key metabolites: Spermine (left), mid-Spermidine (mid), and Glutathione (right).
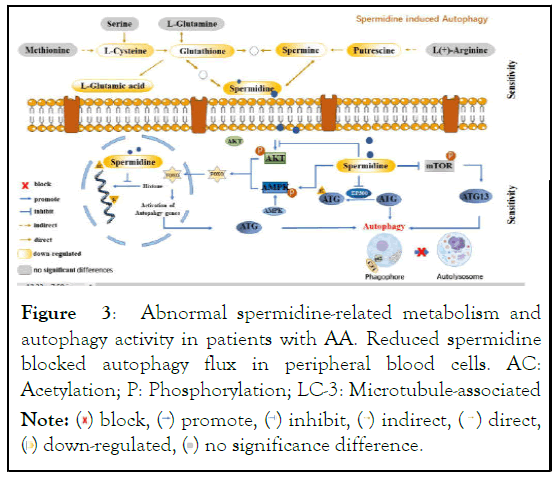
Figure 3: Abnormal spermidine-related metabolism and
autophagy activity in patients with AA. Reduced spermidine
blocked autophagy flux in peripheral blood cells. AC:
Acetylation; P: Phosphorylation; LC-3: Microtubule-associated.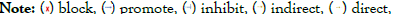
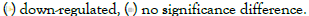
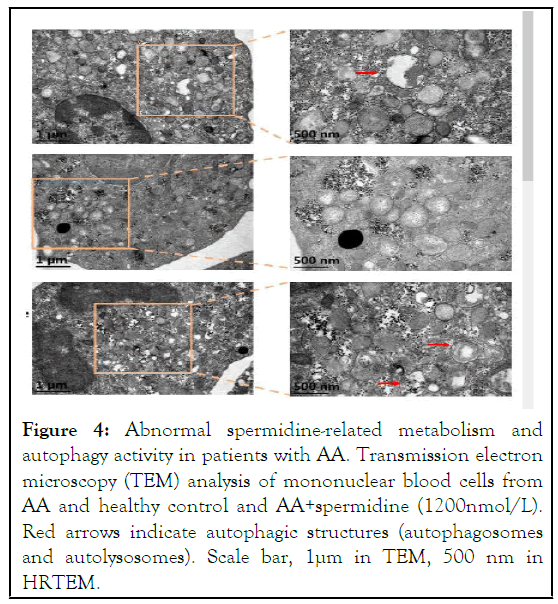
Figure 4: Abnormal spermidine-related metabolism and autophagy activity in patients with AA. Transmission electron microscopy (TEM) analysis of mononuclear blood cells from AA and healthy control and AA+spermidine (1200nmol/L). Red arrows indicate autophagic structures (autophagosomes and autolysosomes). Scale bar, 1μm in TEM, 500 nm in HRTEM.
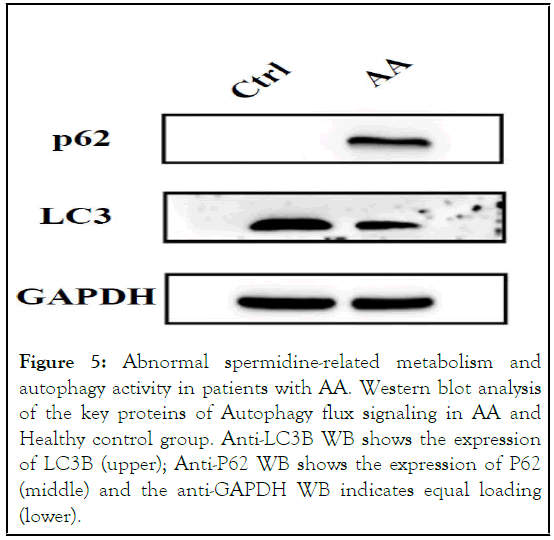
Figure 5: Abnormal spermidine-related metabolism and autophagy activity in patients with AA. Western blot analysis of the key proteins of Autophagy flux signaling in AA and Healthy control group. Anti-LC3B WB shows the expression of LC3B (upper); Anti-P62 WB shows the expression of P62 (middle) and the anti-GAPDH WB indicates equal loading (lower).
protein light chain 3; ATG: Autophagy-related gene or protein; FOXO: Forkhead box transcription factors.
To gain further mechanistic insights, we performed scRNA-seq of bone marrow samples derived from AA patients (n=3)and healthy donors (n=4). After preprocessing with proper experimental controls (Figure S1A), we identified four cell clusters, including HSPCs, B cells, T/NK cells, and myeloid cells (Figure 7 and S1B). These cell clusters were determined by celltype- specific marker genes that were differentially expressed (Figure S1C and S1D). However, there was no difference in the proportion of four cell clusters between healthy donors and AA patients (Figure S1E and S1F). To determine the autophagy level of each cell cluster, we further examined autophagic activity based on the expression of 1,411 human autophagy-related genes retrieved from the Human Autophagy Database. Compared with the healthy donors, we found that some T/NK cell sub-clusters in AA patients have enhanced autophagic activity (Figure S1G). To decipher the characteristics of these enhanced autophagic T/ NK cell populations, we re-clustered 7435 T/NK lymphocytes and identified 11 distinct T/NK cell clusters containing cells from AA patients and healthy donors (Figure S2A). Notably, these enhanced autophagic T cells belonged to Th1, CTL, and Tem subsets. Moreover, these subsets have a significantly higher proportion in AA patients (all P<0.05) (Figure S2B). In terms of T cell function pathways, we found Notch, Jak/STAT, TNF-α, hypoxia, Kras, p53, and unfolded protein signal pathways were upregulated in these three sub-clusters of AA patients (Figure S2D) based on the results of scRNA-seq, all of these signaling pathways being associated with enhanced immunoreactivity. No apparent pathway changes were found for the remaining T cell subsets (Figure S2C). These results suggested that the autophagy activity in Th1, CTL, and Tem is increased, which leads to enhanced T cell immunoreactivity.
Finally, we investigated the mechanisms of enhanced autophagy activity in Th1, CTL, and Tem. It has been reported that overexpression of Spermidine/Spermine N1-acetyltransferase 1 (Sat-1), the rate-limiting enzyme in polyamine catabolism, in human cell lines leads to a rapid depletion of spermidine and spermine [14]. Therefore, we speculated that overexpression of Sat-1 in Th1, CTL, and Tem cell subsets may decrease spermidine levels in AA patients. Indeed, compared to the other bone marrow cells, we found that Th1, CTL, and Tem cell subsets exhibited high Sat-1 transcript levels (all P<0.05) (Figure 6). Moreover, the proportion of this three Sat-1 (+) cell subsets in AA patients was higher than that in normal donors (Figure 7). In addition, based on autophagic activity (Figure.S1G), we classified the myeloid cells into seven sub-clusters (Figure S3A and S3B); however, no overt difference in autophagic activity and proportion between the seven subclusters was found (Figure S3C and S3D), nor was a difference in Sat-1 expression level observed (Figure S3E). These data provide the evidence that hyperactive Th1, CTL, and Tem cells in AA patients overconsumed spermidine through overexpression of Sat-1 [15].
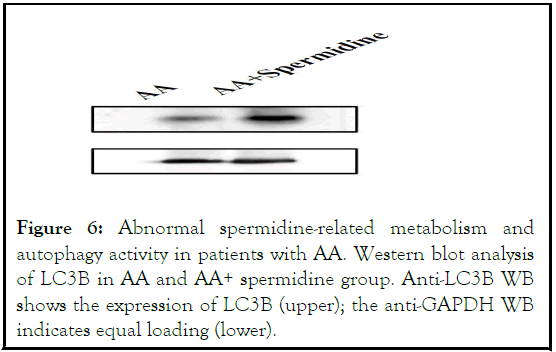

Figure 6: Abnormal spermidine-related metabolism and autophagy activity in patients with AA. Western blot analysis of LC3B in AA and AA+ spermidine group. Anti-LC3B WB shows the expression of LC3B (upper); the anti-GAPDH WB indicates equal loading (lower).
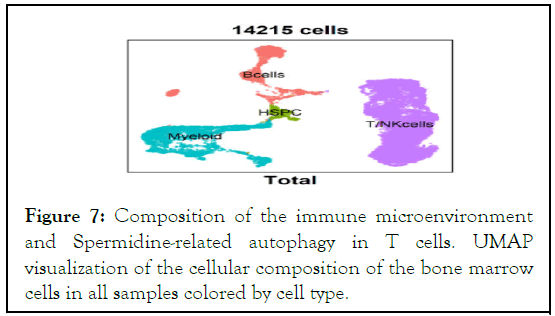
Figure 7: Composition of the immune microenvironment and Spermidine-related autophagy in T cells. UMAP visualization of the cellular composition of the bone marrow cells in all samples colored by cell type.
In this study, we employed a quantitative metabolomics approach to characterize altered metabolic pathways in patients with AA and explored the potential crosstalk between the bone marrow microenvironment and hematopoiesis. To the best of our knowledge, this was the first time that the presence of impaired spermidine metabolism in AA has been discovered. The link of dysregulation of spermidine metabolism to AA allows us to theorize that the level of polyamines may be used as a diagnostic biomarker for AA. Furthermore, we identified three subsets of T cells, including Th1, CTL, and Tem, that were significantly increased in AA patient samples. These three subsets of T cells showed a common feature - an upregulation of the Sat-1 transcription level - which may lead to a rapid depletion of spermidine and spermine. The excessive depletion of spermine in the bone marrow microenvironment could further mediate a decrease in autophagy levels in HSPCs, affecting normal hematopoiesis in AA patients. However, the exact cause of Sat-1 upregulation in AA patients is currently unknown and warrants further investigation. Our data collectively identified that, Th1, CTL, and Tem, key effector cells in AA immune injury, are potential targets for AA therapy, and this underlying mechanism discovered in this study could be uncharted territory for the development of new medications. Based on our above findings, we speculated that targeting Sat-1 may be an effective, safe and viable approach to treating AA.
Jinqi Huang, Ruiqing Zhou, Jian Li, Guo Fu, Kefeng Wu and Qinwei Chen designed the study; Changmei Lin, Yuchan You, Jie Long, Liang Liang, Juan Xia, Yang Chen, Sijie Wang performed and analyzed the experiments; Yuming Zhang, Wenying Luo, Shunqing Wang contributed to patient clinical care and data collection; Jinqi Huang, Qing Li and Qiyuan Li provided scientific advice and supervision; Jinqi Huang, Jian Li, Qinwei Chen, Changmei Lin, Yuchan You and Guo Fu wrotethe paper; and all authors read and approved the final version of the manuscript.
We thank the patient, and his relatives, for his participation in this study, Shujian Chen, Ph.D., for providing technical support, and Mei Fei, Ph.D. for writing and editorial assistance.
This work was primarily supported by the Scientific Research Starting Foundation for Outstanding talent (10101Z20200006), Guangdong Medical University (GDMU). Shunqing Wang and Ruiqing Zhou were supported by the National Natural Science Foundation of China (81600147) and the Key research and development plan of Guangdong Province (2019B020236004). Kefeng Wu and Changmei Lin were supported by the Special Support Project for Southern Marine Science and Engineering Guangdong Laboratory (ZJW-2019-007) and the Discipline construction project of Guangdong Medical University (4SG21009G). Qing Li was supported by the Natural Science Foundation of Fujian Province (No. 2019J01009).
Ethics approval and consent to participate
This study received ethics board approval at Affiliated Hospital of Guangdong Medical University (GDMU).
We obtained the patient’s signed written consent form for the publication of the current research report.
The authors declare that they have no competing interests.
[Crossref][Google Scholar][PubMed].
[Crossref][Google Scholar][PubMed].
[Crossref][Google Scholar][PubMed].
[Crossref][Google Scholar][PubMed].
[Crossref][Google Scholar][PubMed].
[Crossref][Google Scholar][PubMed].
[Crossref][Google Scholar][PubMed].
[Crossref][Google Scholar][PubMed].
[Crossref][Google Scholar][PubMed].
[Crossref][Google Scholar][PubMed].
[Crossref][Google Scholar][PubMed].
Citation: Huanh J, Zhang Y, Xia J, Yang C, Wang S, Liang L, et al. (2022) Spermidine Related Autophagy Flux Deficiency is a Novel Pathogenesis of Aplastic Anemia. J Clin Cell Immunol. 13:669.
Received: 21-Jun-2022, Manuscript No. JCCI-22-18012; Editor assigned: 24-Jun-2022, Pre QC No. JCCI-22-18012 ; Reviewed: 08-Jul-2022, QC No. JCCI-22-18012; Revised: 15-Jul-2022, Manuscript No. JCCI-22-18012; Published: 22-Jul-2022 , DOI: 10.35248/2155-9899.22.13.669
Copyright: © 2022 Huang J, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.