Fisheries and Aquaculture Journal
Open Access
ISSN: 2150-3508
ISSN: 2150-3508
Research Article - (2020)Volume 11, Issue 1
Heritabilities, as well as genetic and phenotypic correlations, were estimated for body weights (BW), body lengths (BL) and body heights (BH) in large yellow croaker, Larimichthys crocea. By crossing the Mindong and Daiqu strain, 60 fullsib families (offspring of 32 males and 60 females) were generated and reared separately. Environmental conditions of each family were standardised, and the body traits were recorded at six months of age. Heritabilities for the body traits were medium to high: 0.31 ± 0.06 for BW, 0.33 ± 0.06 for BL and 0.41 ± 0.07 for BH. The correlations among the three growth traits were positive and high in all cases, with the phenotypic correlations ranging from 0.84 to 0.91 and the genetic correlations ranging from 0.74 to 0.95. The results indicate that the growth traits of juvenile large yellow croaker could be improved efficiently by selection in a future breeding program.
Large yellow croaker; Daiqu and Mindong strains; Growth traits; Genetic parameters
The marine fish species large yellow croaker (Larimichthys crocea) is occurring in the coastal regions of East Asia and are generally found in temperate seawaters (18-25°C). The wild populations have been severely depleted due to overfishing and environment deterioration [1]. The culturing of the species has been successful since the late 1980s, and it is now one of the most important marine aquaculture fish species in China, with a total production of about 180,000 tons in 2017; accounting for more than 12% of the marine fish aquaculture production [2]. Large yellow croaker is a good food resource with excellent nutritional value, providing high quality protein and the healthy omega-3 fatty acids, including eicosapentaenoic acid (EPA) and docosahexaenoic acid (DHA). The swim bladders are also a popular functional food [3], and the market demand for high quality large yellow croaker is increasing. Since the 1980s, large yellow croaker has been cultured and to some extent been subject to phenotypic selection for increased growth, but without pedigree information and control of inbreeding.
Genetic diversity of cultured croaker has thus significantly been reduced compared to the wild populations [4], and the quality has declined for economically important traits, especially growth and disease resistance. An efficient breeding program for large yellow croaker is thus required.
At the beginning of most fish breeding programs, growth is often targeted as the main trait, since it is easy to record and increased growth rate generally improves the profitability of the production. And considerable selection potential for growth has been demonstrated in selective breeding programs of several important aquaculture fish species. For example in Atlantic salmon (Salmo salar), the genetic change per generation was up to 15% in the first generations of the Norwegian selective breeding program [5]; and correspondingly, sustained genetic gain for growth was also high in Nile tilapia (Oreochromis niloticus), with 10–15% per generation in the genetically improved farmed tilapia (GIFT) project [6]. Compared to farm animals, realised genetic responses for increased growth rate are larger in aquatic species [7]. The economic potential of starting a family-based breeding program for increased growth in large yellow croaker should thus be considerable.
Planning of optimal selective breeding requires knowledge of heritabilities and genetic correlations, which are crucial to calculate unbiased breeding values and to predict expected genetic progress. To date, three studies of the genetic parameters for growth traits in large yellow croaker have been reported, but based on the Mindong strain only and with relatively small sample sizes [8-10]. There also exists another geographically and genetically different strain known as the Daiqu strain in the north region of East China Sea [1], whose genetic diversity and growth rate were found to be higher than the Mindong strain [11,12]. Currently, no heritabilities or genetic correlations for growth traits based on crossing of the two strains have been reported. The specific aim of this study was to obtain estimates of these genetic parameters for BW, BL and BH based on crossing individuals from the two strains.
Broodstock fish and design of experiment
Ninety sexually mature fish of Daiqu strain were selected from approximate 1000 individuals at an aquaculture farm in Xiangshan, Zhejiang province (29°5'N, 121°8'E) , and correspondingly, ninety sexually mature fish of Mindong strain were selected from approximate 1000 individuals at another farm in Fuding, Fujian province (27°3'N, 120°25'E). Both strains were sampled with a sex ratio of 2:1 (♀:♂). In March 2014, the broodstock of both strains were transported to the Zhejiang Dahaiyang Science and Technology Co. Ltd. at a sea-site location in Mazhan, Cangnan, Zhejiang province. The mean body weight was 670 ± 123g (mean ± SD) for the Daiqu strain and 500 ± 108g for the Mindong strains. All fish were dorsally tagged with Passive Integrated Transponder (PIT) tags (12 mm × 2.12 mm and 0.09 g). The two strains were kept in two separate tanks (6 m × 8 m × 1.2 m) and fed twice a day with minced fresh mackerels mixed with complex vitamins and mineral additives at a ratio of 5% of body mass per day.
One month later, the brood fish were injected with luteinising hormone-releasing hormone A3 (LHRH A3), with a dose of 2 μg/kilo for females and 1 μg/kilo for males. This resulted in an observed spawning time in the range of 30-34 h post injection [13], when milts and eggs were collected by hand-stripping. A nested design was applied for mating: one Mindong male mated with two Daiqu females, or one Daiqu male mated with two Mindong females. Finally, 32 sires (14 from Daiqu and 18 from Mindong) and 60 dams (27 from Daiqu and 33 from Mindong) produced 60 crossbred fullsib families from the two strains, i.e. comprising 27 fullsib families from Daiqu♂ × Mindong♀-cross and 33 fullsib families from Mindong♂ × Daiqu♀-cross [13].
The culturing of fullsib families
After fertilisation, a standardised quantity of 20 ± 0.2 g floating eggs per family were transferred to separate 1 m3 breeding cylinders for incubation. After hatching, the larvae were fed with rotifers, artemia and copepods, successively. The seawater condition was standardised, with salinity at 24.2 ± 0.1%, temperature at 23.7 ± 0.1°C, dissolved oxygen level at 5-7 mg/L and pH at approximately 8. The water exchange rate was set to 30% per day.
After 50 days, when the average total length of fingerlings was 5 cm, about 1000 randomly sampled fingerlings per fullsib family were transferred to separate small net cages (1 m × 1 m × 2 m) in the sea for on-growing until tagging. They were fed twice a day with commercial fish pellets at a ratio of 5% of body mass (52-55% fish protein powder, 2% fish oil, Shanghai Nonghao Co. Ltd.).
Data collection
In September 2014, when the fish were 6 months old, a random sample of 100 individuals per family (total 6000) were anesthetised with MS-222 (20-30 ppm) after starvation for two days, and body weight, body length (from tip of snout to the last vertebra) and body height (in front of the first ray of dorsal fin from the dorsal margin of the body to the ventral margin) were measured (Figure 1). Afterwards, they were tagged with a PIT tag in the abdominal cavity (Figure 2). The measuring and tagging time per fish was about one minute. If the tagged fish did not recover within five minutes, it was replaced by another fish from the same family.
Figure 1: Body measurements taken on large yellow croaker: BL and BH.
Figure 2: PIT tagging fish with hand-held injection device.
The tagged fish were hereafter cultured communally in one larger net cage (3 m × 3 m × 4 m). The fish were intended for further growth and later selection and mating, but due to a devastating typhoon in August the following year, all the tagged fish were lost. The results reported here are thus based on the data recorded at tagging, i.e. at six months of age.
Statistical analysis
The estimations of heritability and correlations between body weight, body length and body height were achieved with a multitrait animal model, using the ASReml software (version 4.1) [14]. The following model was used, in matrix notation:
y = Xb + Z1c + Z2a + e
Where, y is a vector of individual observations, partitioned for each growth trait, b is a vector of fixed effects (i.e. the Daiqu or Mindong strains and 14 recording dates), a is a vector of random additive genetic effects (number of individuals=6000), c is a vector of common environmental effects pertaining to fullsibs (60 families) and e is a vector of individual random error effects. X, Z1 and Z2 are known design matrices assigning observations to levels of b, c and a, respectively. The heritability was calculated as 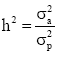 , where
, where 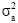 is the additive variance of each trait and
is the additive variance of each trait and 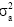 is the phenotypic variance of each trait.
is the phenotypic variance of each trait.
The additive genetic correlations between traits, i and j, was calculated as 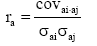 and the phenotypic correlations as
and the phenotypic correlations as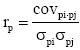 where covi, j is the covariance and σi, σj are standard deviations of the two traits, i.e. for the additive genetic (a) or phenotypic (p) effects, respectively.
where covi, j is the covariance and σi, σj are standard deviations of the two traits, i.e. for the additive genetic (a) or phenotypic (p) effects, respectively.
Growth traits, i.e. BW, BL and BH, of 6000 six-month old individuals from 32 sires and 60 dams were measured at tagging. Descriptive statistics for the three traits are given in Table 1. Twenty-three possible outliers were noted by ASReml for each trait but were still found to be plausible observations after individual checking. The relative variation, quantified by the coefficient of variation (CV), was high for BW but relatively low for BL and BH.
| Mean | SD | CV (%) | |
|---|---|---|---|
| Body weight(g) | 66 | 22 | 33 |
| Body length (cm) | 15.6 | 1.7 | 11 |
| Body height(cm) | 4.2 | 0.6 | 13 |
Table 1: Means, standard deviations (SD) and coefficients of variation (CV) of body weight, body length and body height at 6 months’ age of the F1 generation (N=6000).
Genetic parameters for the three growth traits are presented in Table 2. Heritabilities for all the three growth traits were high, ranging from 0.31 (BW) to 0.41 (BH). The phenotypic and genetic correlations among these growth traits were also relatively high. The highest phenotypic (0.91) and genetic correlation (0.95) was between BW and BL, while the lowest correlations were between BL and BH (rp=0.84, rg=0.74). The genetic correlation between BW and BL was higher than the phenotypic correlation, while the opposite was found for the other traits. The fullsib family variance component, associated with the common environment effect, was not estimable in the present dataset and was set to zero.
| Body weight | Body length | Body height | |
|---|---|---|---|
| Body weight | 0.31 ± 0.06 | 0.95 ± 0.02 | 0.79 ± 0.06 |
| Body length | 0.91 ± 0.004 | 0.34 ± 0.06 | 0.74 ± 0.07 |
| Body height | 0.87 ± 0.01 | 0.84 ± 0.01 | 0.41 ± 0.07 |
Table 2: Heritability (diagonal, in bold), genetic (above diagonal) and phenotypic (below diagonal) correlations with standard errors (±SE) between body weight, body length and body height.
The heritabilities for the measured growth traits of juvenile large yellow croaker ranged from 0.31 to 0.41 in the present study, suggesting that rapid genetic gains could be achieved by BLUP selection for these traits in juvenile fish. Generally, selection should be made close to market size of the fish, but due to loss of all tagged fish in the typhoon season, heritability for growth at harvest was not available in this study. Liu et al. [8] estimated heritabilities for growth traits of 20 month old fish in the range of 0.09-0.19 based on 599 individuals, but based only on two fullsib/halfsib groups (4 males × 4 females and 4 males × 3 females); and in another study [9] in 13-month old fish the estimates ranged from 0.02 to 0.36, based on 959 offspring from only 10 dams and 15 sires; whereas Dong et al. [10] estimated heritabilities for BW and BL to be 0.60 and 0.59, respectively, using genomic prediction based on 500 two-year old fish. Heritabilities for growth traits will often vary with age in fish; for instance, higher heritabilities for body weight and total length were found at 270 days post hatch (dph) than at 90 dph in Asian seabass [15] and a similar trend was found in rainbow trout [16]. It is still uncertain whether heritabilities for growth traits increase or decrease with age in large yellow croaker.
Due to separate rearing of fullsib families before tagging, any common environmental effects could be confounded with additive genetic effect in the statistical analysis, and heritability estimates may thus be inflated in this study. However, when the common environmental effect was fitted in the model, it was not estimable, and ASReml automatically set the effect to zero in order to keep the solutions within the parameter space. Unfortunately, the hierarchical design used in this study is not well suited for separation of genetic and environmental effects pertaining to the same fullsib family. Still, the results may also indicate that the effect was indeed not large, which could be due to proper standardisation of the fullsib environments during the pre-tagging period, since the juvenile fish were reared in the sea most of the time, which to a large extent implied sharing equal seawater quality. Although common environmental effects have been shown to be considerable in many aquaculture species, ranging from 5% to 20% [17,18], a recent study also showed that this effect may to a large degree be due to maternal effects [19], as it was found that the effect accounted for as much as ~10% of the overall variance, despite eliminating all common environmental effects by use of a marker based tagging system, i.e. no separate rearing of families were required. The maternal effects may thus very well be smaller in marine species, like large yellow croaker, having smaller eggs than freshwater species.
Heterosis and reciprocal effect are other factors that may inflate the heritability estimates, and these effects could occur in the present design as we were doing crosses between the Mindong and Daiqu strains. Both effects are important to determine the optimum utilisation and setup for the different strain crosses in an eventual cross-breeding program. In tilapia, significant heterosis and reciprocal effects of growth traits have been observed in diallel crosses, e.g. 0-12% in Bentsen et al. and 6-12% in Said and Mekkawy [20,21]. However, we were not able to estimate any heterosis or reciprocal effects in our design, as only cross-strain mating and no pure-strain mating were performed. As mentioned above, the purpose here was to form a new mixed base population, and no crossbreeding scheme is planned in the future. Handling and mating of yellow croakers is still exceptional difficult, and even a simplified design, as in the present study, is a challenging task.
Genetic and phenotypic correlations among the growth traits of the fish were found all positive and high, especially between BW and BL. In an earlier study, genetic correlations among the same three traits were even higher than the present study, all above 0.95 [8]. Similar high genetic correlations have also been reported in other fish species; for instance in Nile tilapia (Oreochromis niloticus), the genetic correlation between body weight and body length at harvest was close to unity [22] , and in six months old coho salmon (Oncorhynchus kisutch) it was 0.98 ± 0.06 [23]. This may indicate that BW and BL are largely controlled by pleiotropic genes and selection could thus be conducted on either of the two traits with little difference in selection response. Considering easier measurement and higher heritability and CV for body weight, it is suggested that body weight should be used as the selection criterion.
The present genetic parameter estimates for large yellow croaker were based on records of six-month-old fish at tagging. However, growth until harvest is a more economically important trait for any breeding program. The validity of these genetic parameter estimates for growth at harvest time is uncertain, but in other cultured fish species, high positive genetic correlations between growth intervals have been reported. For example, the average genetic correlation for weight at various ages was 0.7 in European sea bass and 0.93 in red drum [24,25], although a study in rainbow trout showed that for distant life periods it can be as low as 0.24 [26]. The genetic parameters estimated for juvenile fish in this study could still be useful predictors if pre-selection is performed instead of selection at harvest in yellow croaker, which will be the case if for instance a modern DNA-based tagging system is used. But the genetic correlation between the two growth stages is still needed and should be verified based on more experiments.
In the study, the highest coefficient of variation (CV) was found for BW, which is consistent with a previous study for 20 months old fish [8]. The average tagging weight was 66 ± 22 g for BW, which was larger than usual for other fish species at tagging, using 11 mm PIT-tags. For instance, in rohu carp (Labeo rohita), fingerlings at 8-15 g were found to be suitable for tagging with similar PIT tags [27]. The main reason for delayed tagging in our study was that the experiment experienced typhoon season during summer and the cages used were not storm-resistant.
In conclusion, the present results show that there is a potential to perform family selection for BW, BL and BH at an early age of large yellow croaker, as medium to high heritabilities were found for these traits at six months of age, and among these three traits, BW should be used as the selection criterion, since both the heritability and CV is highest for this trait. Future work and resources are needed to establish a full operational and secured breeding program for this fish species.
This study was approved by the Institutional Animal Care and Use Committee (IACUC) of Zhejiang Ocean University, Zhoushan, China, and this research was financially supported by the Open Foundation from Marine Sciences in the Most Important Subjects of Zhejiang (No.20130204). The authors are also thankful to the help of Zhejiang Dahaiyang Science and Technology Co., Ltd.
Citation: Yu X, Ådnøy T, Lv Z, Wu C, Gjøen HM (2020) Phenotypic and Genetic Parameter Estimation for Growth Traits in Juvenile Large Yellow Croaker (Larimichthys crocea). Fish Aqua J 11:274. doi: 10.35248/2150-3508.20.11.274
Received: 09-Jan-2020 Accepted: 15-Feb-2020 Published: 22-Feb-2020 , DOI: 10.35248/2150-3508.20.11.274
Copyright: © 2020 Yu X, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.