Rheumatology: Current Research
Open Access
ISSN: 2161-1149 (Printed)
ISSN: 2161-1149 (Printed)
Research Article - (2022)Volume 12, Issue 5
Introduction: Herpes Zoster (HZ) is known as a side effect of using biologics in Rheumatoid Arthritis (RA). Incidence of this side effect may be different depending on genetic factors because susceptibility to HZ infection varies by race. Here, we analyzed the statistical relationships of whole genome Single Nucleotide Polymorphisms (SNPs) with HZ infection in biologics-treated RA patients.
Materials and methods: The subjects were 321 Japanese female RA patients (including 56 herpes virus infected patients) using biologics. The relationships of 302,814 SNPs with HZ infection were analyzed using case-control analyses by the Chi-square test. We picked up SNPs (P<10-8) significantly associated with HZ infection. Then, the effects of the SNPs were analyzed by using a multivariate logistic regression analysis adjusted for the age of first biologics administration, disease activity, corticosteroid use and methotrexate use.
Results: Rs10774580 located in 2’-5’-oligoadenylate synthetase like gene (OASL) was significantly associated with HZ infection. The minor allele homozygous carrier was positively associated with HZ infection in multivariate analysis.
Conclusion: We for the first time showed a significant relationship between a genetic factor and HZ infection among RA patients. Rs10774580 may be one of the biomarkers for HZ infection.
Herpes zoster; Rheumatoid arthritis; Biologics; Infection; Genome-Wide Association Study (GWAS); Single Nucleotide Polymorphism (SNP)
Rheumatoid Arthritis (RA) is a progressive autoimmune disease well defined by widely accepted symptoms such as chronic joint inflammation and structural damage [1]. In treatment for RA at present, using biological agents such as Tumor Necrosis Factor (TNF), Interleukin-6 (IL-6) and Cytotoxic T-Lymphocyte-Associated protein 4 (CTLA-4) blockades is extremely useful because these agents specifically inhibit immune responses and inflammation. On the other hand, because these agents are immunosuppressant, infectious diseases become a significant problem in treatment for RA.
Herpes Zoster (HZ) is one of the most common viral infections in treatment for RA with immunosuppressants such as biological agents. In fact, it has been reported that RA patients have a higher risk of HZ compared with the general population [2-4]. Therefore, a number of studies analyzed relationships of the incidence of HZ with various possible risk factors. As the result, aging, high disease activity corticosteroid, methotrexate (MTX) and biological agents were reported as risk factors for HZ [5]. These studies, however, have not taken genetic factors into consideration. On the other hand, several studies have identified genetic loci associated with onset of RA [6]. It has also been reported that the effectiveness of biologic agents can be predicted by a combination of Single Nucleotide Polymorphisms (SNPs) [7]. These studies used Genome- Wide Association Study (GWAS) in order to identify the genetic factors. Thus, conducting GWAS is thought to be valuable in order to identify unknown genetic factors associated with HZ in RA.
In this study, in order to identify SNPs associated with HZ infection, we analyzed the statistical relationships of whole genome SNPs with HZ infection among biological agents-treated RA patients.
Patients
We recruited 321 Japanese female RA patients receiving treatment with biological disease-modifying antirheumatic drugs (bDMARDs). We researched whether or not each patient was infected with HZ within 6 years from the date of first administration of bDMARDs. All patients had been receiving treatment with bDMARDs for the 6 years. In addition, they hadn’t gotten the vaccination against HZ before we recruited them. They also hadn’t gotten the vaccination during their follow-up period. Written informed consent to participate in this study was obtained from each patient. This study was approved by the ethical committee for analytical research on the human genome of the Matsubara Mayflower Hospital. All methods were performed in accordance with relevant guidelines and regulations.
Assessment of disease activity
The Disease Activity Score with 28 joints using C-reactive protein (DAS28-CRP) was used in order to assess disease activity for each patient timed with the start of bDMARDs.
Genome-wide SNP genotyping
The patients’ whole blood samples were used for DNA extraction at Mitsubishi BCL Inc. Genome wide SNP genotyping were performed at deCode genetics Inc. (Reykjavic, Iceland) using Illumina HumanHap300K chip technology (Illumina Corp., San Diego, CA, USA). After genotyping, 302,814 of 317,503 SNPs excluded SNPs with call rates <90% and minor allele frequency <1 % were used in the case-control analysis described below.
Statistical analysis
We used Chi-square test to compare the frequencies of the age of first bDMARDs administration, disease activity, corticosteroid use and MTX use between the HZ non-infected and the infected patients. We also used Mann-Whitney U test to compare the median values of the age of first bDMARDs administration, DAS28-CRP score, dose of corticosteroid and dose of MTX between the HZ non-infected and the infected patients. We used case-control analysis to analyze the relationships of 302,814 SNPs with the susceptibility to HZ infection by Chi-square test using SVS 8.1.1 (Golden Helix Inc.). After case-control analyses, we picked up SNPs significantly associated with HZ infection. The genotype frequency was compared between the HZ non-infected and the infected patients using Bonferroni-corrected Chi-square test. Univariate and multivariate logistic regression analyses were used to analyze the effects of the SNP, the age of first bDMARDs administration, disease activity, corticosteroid use and MTX use on the risk for HZ infection. The logistic regression analyses were carried out using EZR (Saitama Medical Center, Jichi Medical University, Saitama, Japan) [8]. EZR is a graphical user interface for R (The R Foundation for Statistical Computing, Vienna, Austria, version 2.13.0). P-values <10-8 were considered significant in case- control analysis. P-values <0.05 were also considered significant in the logistic regression analyses.
The basic characteristics of the patients are presented in Table 1. The total patients were 321. The median age of first bDMARDs administration in HZ infected patients was significantly higher than those in HZ non-infected patients. The frequency of higher age (≥ 57 years) of the administration in HZ infected patients was significantly higher than those in HZ non-infected patients. There were no significant differences in the frequency of disease activity, corticosteroid use and MTX use between the HZ non-infected and infected patients. There were no significant differences in the median of DAS28-CRP score, corticosteroid dose and MTX dose between them.
| Variables | All patients | HZ non-infected | HZ infected |
|---|---|---|---|
| Number of patients (%) | 321 | 265 (82.6) | 56 (17.4) |
| Age of first administration (years) | 56 (48,65) | 57 (47,65) | 61 (55,67)a |
| Higher age (≥ 57 years) (%) | 159 (50.0) | 122 (46.0) | 37 (66.1)a |
| Onset age of HZ infection (years) | ̶ | ̶ | 69 (64, 75) |
| Disease activity (DAS28-CRP score) | 4.9 (3.9,5.7) | 4.9 (3.9,5.7) | 5.0 (4.0,5.9) |
| High activity (>4.1) (%) | 196 (61.1) | 159 (60.0) | 37 (66.1) |
| Unknown (%) | 23 (7.2) | 17 (6.4) | 6 (10.7) |
| Corticosteroid use (%) | 172 (58.1) | 136 (56) | 36 (67.9) |
| Unknown (%) | 25 (7.8) | 22 (8.3) | 3 (5.4) |
| dose (mg/day) | 5.0 (3.0, 5.0) | 5.0 (3.0, 5.0) | 4.5 (3.0, 5.0) |
| MTX use (%) | 179 (55.8) | 140 (58.1) | 39 (73.6) |
| Unknown (%) | 27 (8.4) | 24 (9.1) | 3 (5.4) |
| Dose (mg/week) | 8.0 (6.0, 8.0) | 8.0 (6.0, 8.0) | 8.0 (6.0, 8.0) |
| Abbreviations: HZ, herpes zoster. Values are median (IQR), number of the patients Note: aP-value<0.01 vs. HZ non-infected. |
|||
Only one SNP was identified that was significantly associated with HZ infection (Figure 1). The SNP was rs10774580 located in 2’-5’-oligoadenylate synthetase like gene (OASL).
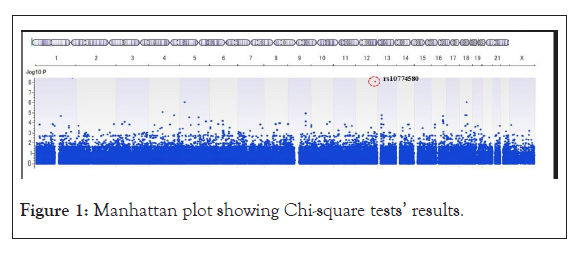
Figure 1: Manhattan plot showing Chi-square tests’ results.
Table 2 presents the frequency of rs10774580. In HZ infected patients, the frequency of minor allele homozygous was significantly higher than that of major allele homozygous and heterozygous.
| OASL genotype (%) | All patients | HZ non-infected | HZ infected |
|---|---|---|---|
| Major allele homozygous | 186 (58.0) | 160 (60.4) | 26 (46.4) |
| Heterozygous | 122 (38.0) | 102 (38.5) | 20 (35.7) |
| Minor allele homozygous | 13 (4.0) | 3 (1.1) | 10 (17.9)a |
| Abbreviation: HZ: Herpes Zoster. Note: aP value<0.01 vs. HZ non-infected. |
|||
Table 3 presents the relationships of OASL genotype, the age of first bDMARDs administration, disease activity, corticosteroid use and MTX use with HZ infection. The minor allele homozygous of rs10774580 was positively associated with HZ infection. Adjusted OR for the minor allele homozygous was 39.2 (95 % CI 4.6−331). There were no significant relationships of the age of first bDMARDs administration, disease activity, corticosteroid use and MTX use with HZ infection.
| Variables (high-risk group) | Univariate | Multivariatea | ||
|---|---|---|---|---|
| OR (95% CI) | P-value | OR | P-value | |
| OASL genotype (minor allele homozygous) | 19.0 (5.0-71.6) | <0.01 | 39.2 (4.6-331) | <0.01 |
| Age of first bDMARDs administration (≥ 57 years) | 2.1 (1.1-3.8) | 0.021 | 1.6 (0.83-3.3) | 0.16 |
| Disease activity (high activity) | 1.6 (0.81-3.2) | 0.18 | 1.1 (0.50-2.3) | 0.88 |
| Corticosteroid use | 1.7 (0.89-3.1) | 0.11 | 1.6 (0.75-3.6) | 0.22 |
| MTX use | 2.0 (1.0-3.9) | 0.039 | 1.5 (0.67-3.3) | 0.33 |
| Abbreviation: OR: Odds Ratio. Note: aAdjusted for all variables |
||||
To our knowledge, our study is the first to analyze the relationships of whole genome SNPs with HZ infection among bDMARDS- treated RA patients and to identify a SNP as one of the biomarkers for HZ infection.
It has been reported that aging, high disease activity, corticosteroid use and MTX use are risk factors for HZ infection in RA patients [5]. In fact, age at first administration of bDMARDs in HZ infected patients was significantly higher than that in HZ non- infected patients in our study. In addition, it has been reported that susceptibility to HZ infection in RA varies by race [9]. In fact, Japanese and Taiwanese have a higher risk compared to Americans and Europeans [10]. In this regard, genetic background may also affect the susceptibility. However, because there are no studies taking into account genetic polymorphisms such as SNPs, it is likely that genetic polymorphisms associated with the susceptibility were overlooked. Therefore, we conducted GWAS which is powerful tool to collectively identify SNPs associated with susceptibility.
In our study, rs10774580 located in intron region of OASL gene was significantly associated with HZ infection. The minor allele homozygous were positively associated with HZ infection. Human OASL has an antiviral activity against RNA viruses [11,12]. On the other hand, OASL inhibits type I interferon (IFN) induction during DNA virus infection such as herpes simplex, vaccinia and adenovirus [13]. This is because OASL binds to cylic GMP-AMP synthase (cGAS) known as DNA sensor, and inhibits cyclic GMP- AMP (cGAMP) synthesis in cGAS-STING (stimulator of interferon gene) pathway sensing the majority of DNA viruses [13]. Inhibiting IFN induction leads to enhancing DNA virus replication. Therefore, rs10774580 may affect the transcription of OASL because this SNP is intronic variation without amino acid substitution. According to the GTEx database (www.gtexportal.org/home/) [14], rs10774580 has been reported as an expression Quantitative Trait Loci (eQTL) of OASL in whole blood and lung. Therefore, though the difference of susceptibility to HZ infection is likely to reflect the change of expression levels among OASL genotypes, the detailed mechanism is unclear. Thus, the functional analyses are needed with regard to OASL expression among the genotypes.
Interestingly, several previous studies revealed that using Janus Kinase (JAK) inhibitors increased the risk of HZ infection compared to bDMARDS [15,16]. In this regard, it is unclear if rs10774580 is also associated with HZ infection in JAK inhibitors-treated RA patients. Thus, further analyses are needed in JAK inhibitors- treated RA patients.
This study has several limitations. First, rs10774580 was identified by the result of GWAS among Japanese RA patients. It is well known that allele frequencies of most SNPs vary in different ethnic groups. The allele frequency of rs10774580 we identified also varied compared with the allele frequencies of many other ethnic groups reported in the HapMap database (https://www.ncbi.nlm.nih.gov/ snp). However, the frequencies of the SNP in Japanese, American and South Asian are almost the same. Therefore, rs10774580 may be applicable to American and South Asian RA patients as one of the biomarkers. In this regard, further studies are needed in these populations. A second limitation is that this study didn’t take into consideration the number of incidences of HZ in each patient. It is well known that some RA patients repeatedly develop HZ. Therefore, in order to identify the other biomarkers, further studies taking the incidence of HZ into consideration are desired.
This is the first report of a significant association between a genetic factor and HZ infection among bDMARDS-treated RA patients. As the results of GWAS and multivariate logistic regression analysis, we showed that rs10774580 in OASL gene was significantly associated with HZ infection. Therefore, this SNP may be one of the biomarkers for predicting HZ infection among RA patients before using biologics.
Acknowledgements
We would like to thank Dr. Eisuke Shono, Tomomaro Izumihara, Tomomi Tsuru and Motohiro Oribe for offering the patients’ whole blood samples. We also would like to thank Ms. Carol Matsubara for proofreading the manuscript.
Conflicts of interest
The authors had no conflicts of interest.
[Crossref] [Google scholar] [pubmed]
[Crossref] [Google scholar] [pubmed]
[Crossref] [Google scholar] [pubmed]
[Crossref] [Google scholar] [pubmed]
[Crossref] [Google scholar] [pubmed]
[Crossref] [Google scholar] [pubmed]
[Crossref] [Google scholar] [pubmed]
[Crossref] [Google scholar] [pubmed]
[Crossref] [Google scholar] [pubmed]
[Crossref] [Google scholar] [pubmed]
[Crossref] [Google scholar] [pubmed]
[Crossref] [Google scholar] [pubmed]
[Crossref] [Google scholar] [pubmed]
[Crossref] [Google scholar] [pubmed]
[Crossref] [Google scholar] [pubmed]
Citation: Hashimoto M, Funahashi K, Tsumiyama K, Takashima Y, Maeda T, Fukuda K, et al. (2022) OASL rs10774580 SNP is Associated with Herpes Zoster Infection in Rheumatoid Arthritis. J Rheumatol. 12: 312.
Received: 11-Aug-2022, Manuscript No. RCR-22-18817; Editor assigned: 16-Aug-2022, Pre QC No. RCR-22-18817 (PQ); Reviewed: 24-Aug-2022, QC No. RCR-22-18817; Revised: 01-Sep-2022, Manuscript No. RCR-22-18817 (R); Published: 09-Sep-2022 , DOI: 10.35841/2161-1149.22.12.312
Copyright: © 2022 Hashimoto M, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.